



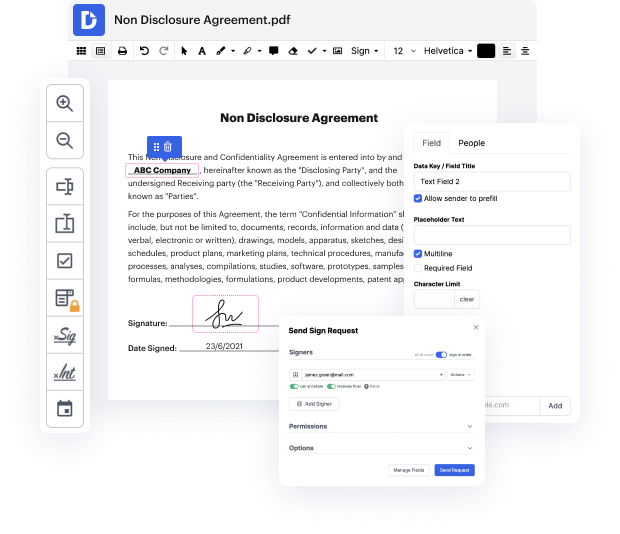
Picking out the perfect file managing solution for your firm might be time-consuming. You have to analyze all nuances of the app you are interested in, compare price plans, and stay vigilant with security standards. Arguably, the ability to work with all formats, including rtf, is essential in considering a platform. DocHub provides an substantial list of features and tools to successfully manage tasks of any difficulty and handle rtf file format. Register a DocHub profile, set up your workspace, and begin working with your files.
DocHub is a thorough all-in-one program that lets you modify your files, eSign them, and make reusable Templates for the most commonly used forms. It offers an intuitive interface and the ability to manage your contracts and agreements in rtf file format in a simplified mode. You do not have to bother about reading countless guides and feeling stressed out because the software is too complex. remove effect in rtf, assign fillable fields to chosen recipients and gather signatures quickly. DocHub is all about potent features for specialists of all backgrounds and needs.
Improve your file generation and approval procedures with DocHub today. Enjoy all this with a free trial version and upgrade your profile when you are ready. Edit your files, produce forms, and learn everything that can be done with DocHub.
Hello from GenomeSpace! In this video step we will attempt to remove batch effects from our gene expression dataset using the ComBat module in GenePattern. From the Modules tab search or browse for the ComBat module. ComBat requires continuous gene expression data such as microarray intensity. Thus, we previously converted our discrete RNA-Seq count data to the continuous log counts-per-million unit using the PreprocessReadCounts module. We access the outputs from the PreprocessReadCounts module under the Jobs tab. Click and drag the gene expression file, in this case GSE63412GSE44229.logcpm.gct, to the input file field. ComBat also requires a sample information file that annotates the samples for the batch covariates as well as covariates of interest. From the GenomeSpace tab, click and drag a sample info file such as GSE63412GSE44229.sample.info.txt to the sample info file field. We will be using the parametric prior method. If the parametric estimates are poor, we can rer
At DocHub, your data security is our priority. We follow HIPAA, SOC2, GDPR, and other standards, so you can work on your documents with confidence.
Learn more



