



Are you having a hard time finding a reliable solution to Annotate Table Letter For Free? DocHub is made to make this or any other process built around documents much easier. It's straightforward to explore, use, and make edits to the document whenever you need it. You can access the core tools for handling document-based workflows, like certifying, adding text, etc., even with a free plan. Moreover, DocHub integrates with different Google Workspace apps as well as services, making file exporting and importing a breeze.
DocHub makes it easier to work on paperwork from wherever you’re. In addition, you no longer need to have to print and scan documents back and forth in order to sign them or send them for signature. All the essential tools are at your disposal! Save time and hassle by executing paperwork in just a few clicks. a go today!
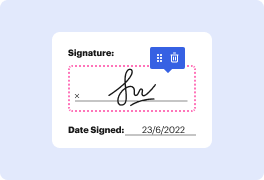
The user has logged into the Gene Expression platform and accessed their saved files. They are working on genes upregulated upon TNF stimulation in Hovex cells, previously converted to a list with identifiers. They find the table with only gene identifiers challenging and plan to add gene descriptions and symbols using the annotate table method. They locate the method in the Analysis tab, input their gene table, and select the human annotation source.
At DocHub, your data security is our priority. We follow HIPAA, SOC2, GDPR, and other standards, so you can work on your documents with confidence.
Learn more



